Reaction Path Averaging
published 2017-02-13
The DNA is one of the best-known biomolecules. Upon electromagnetic radiation, an electron may be released from a nucleobase, resulting in a positive electron hole. The hole may travel across the nucleobases turning the DNA into a sort of nanowire. This is a rather established fact. The shape of the DNA is so classic that it is natural to ask how the DNA distorts upon the electron transfer. With Tom Kubař we've recently attempted to answer. We used a computational approach, which we called Reaction Path Averaging (RPA).
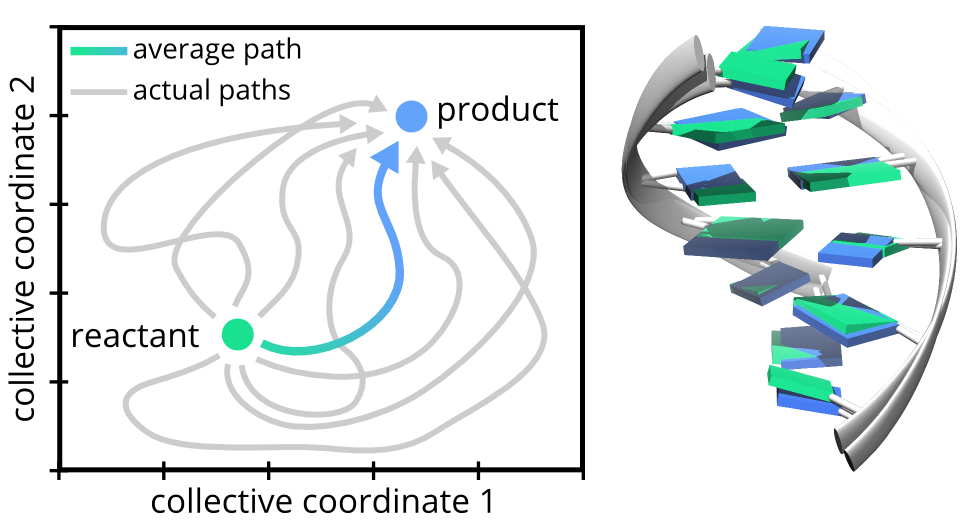
The electron transfer is, in fact, a simple chemical reaction. The "reactant" is a DNA with the charge localized on certain nucleobase, the "product" (of the reaction) is a DNA with the charged localized somewhere else. The transfer between two neighboring nucleobases may happen via a myriad of routes (or reaction paths). By means of molecular dynamics simulations, we generated thousands of such paths and analyzed them.
We've found that the average reaction path is quite simple. Upon electron transfer, the DNA distortions reduce (on average) into the shift of the nucleobases in the direction from/to the minor/major groove. According to our simulation, the relaxation of the DNA structure takes about 1 ns. Is such shaking fast?
During the project, we faced some (funny) technical problems. For instance, we needed to generate an ensemble of initial conformations of the DNA by equilibrium molecular dynamics. It turned out that this generates about 20 GB/hod (= 480 GB/day = 3.4 TB/week), which we really didn't expect. My laptop free disk space would be filled within 90 mins.
