Tutorial: How to Visualize a Molecular Electrostatic Potential with Molekel
published 2017-08-16
The electrostatic potential is a physical quantity that is valuable for understanding intermolecular interactions such as hydrogen or halogen bonding. It is instructive to project the electrostatic potential onto a molecular surface. Such a visualization, also known as Molecular Electrostatic Potential (MEP), can be done in Molekel 4.3. It is a rather old and not-so-intuitive program with poor documentation, so in this tutorial I decided to describe how to get pictures like this:
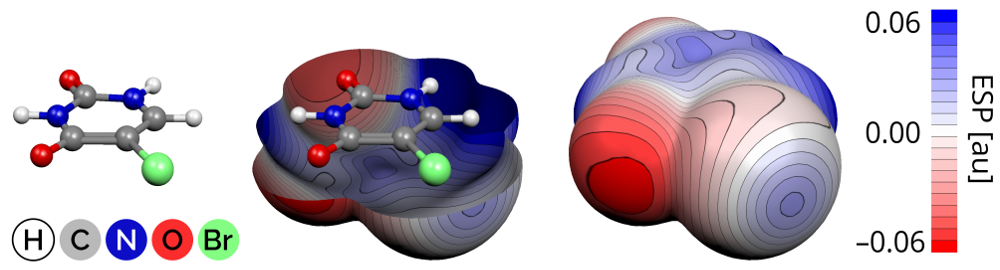
I will assume a working knowledge of an Unix-like operating system and quantum chemistry programs. Luckily, there are tons of documentation of both available online.
We need two sorts of volumetric data: i) electron density for the surface construction and ii) electrostatic potential for the "coloring" of the surface. We obtain them as "cube files" with Gaussian program package. By adding a line with "%chk=mol.chk" into the Gaussian input file, we ask the program to keep the checkpoint file that we subsequently convert into a formatted checkpoint file by formchk.
formchk mol.chk mol.fchk
The mol.fchk serves as input for the cubegen program that generates the desired cube files. We need to run it twice - for the density and the potential separately.
cubegen 0 density=scf mol.fchk mol.dens.cube -3 h
cubegen 0 potential=scf mol.fchk mol.pote.cube -3 h
Now we're ready to move to the Molekel. If we run it without arguments from the terminal, we get a blank black screen (main molekel window) and main control panel. Right click the main window shows the main menu:
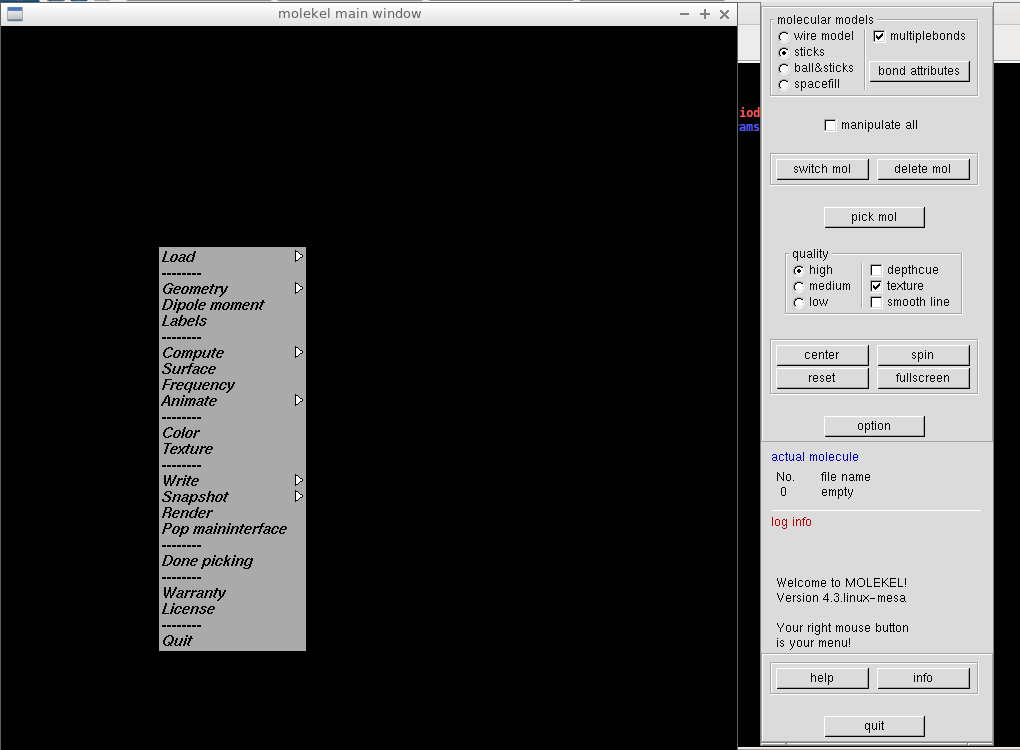
We select "Load/Gaussian cube" and in the following window, we go for the mol.dens.cube. A structure of our molecule should appear. Right click on it shows the main menu, and we select "Surface." A new window opens, where we check "gaussian cube" instead of the default "macu" file type, and load the density cube file again. Molekel's main control panel shows some basic info about the cube file, and in the Surface window, a non-zero V-max value should appear.
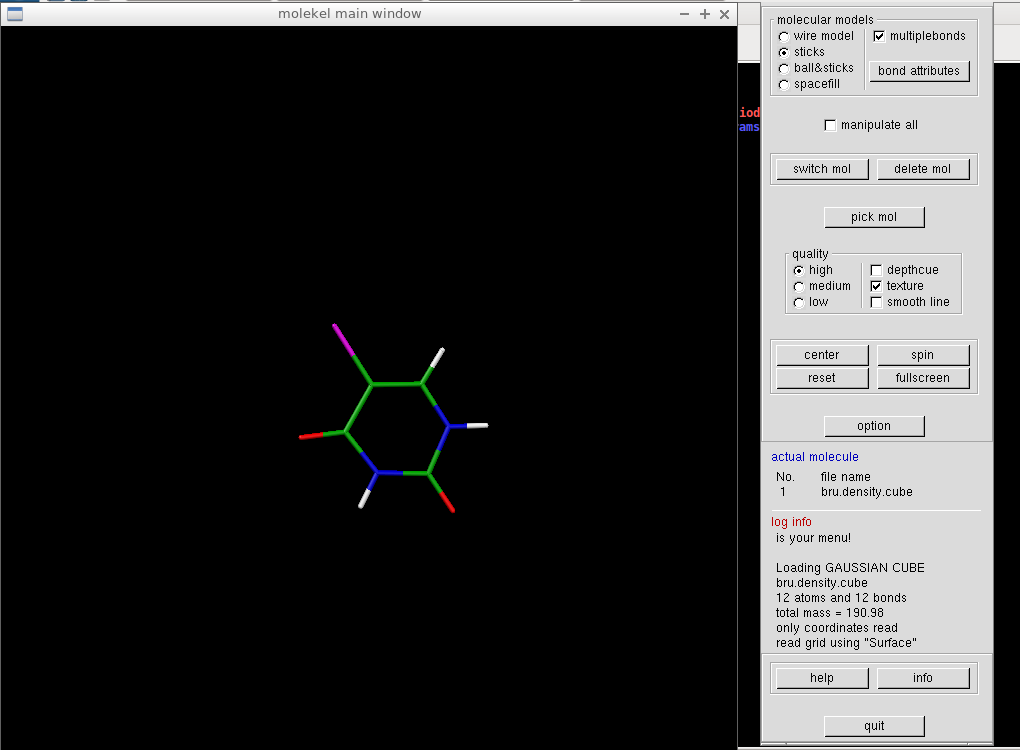
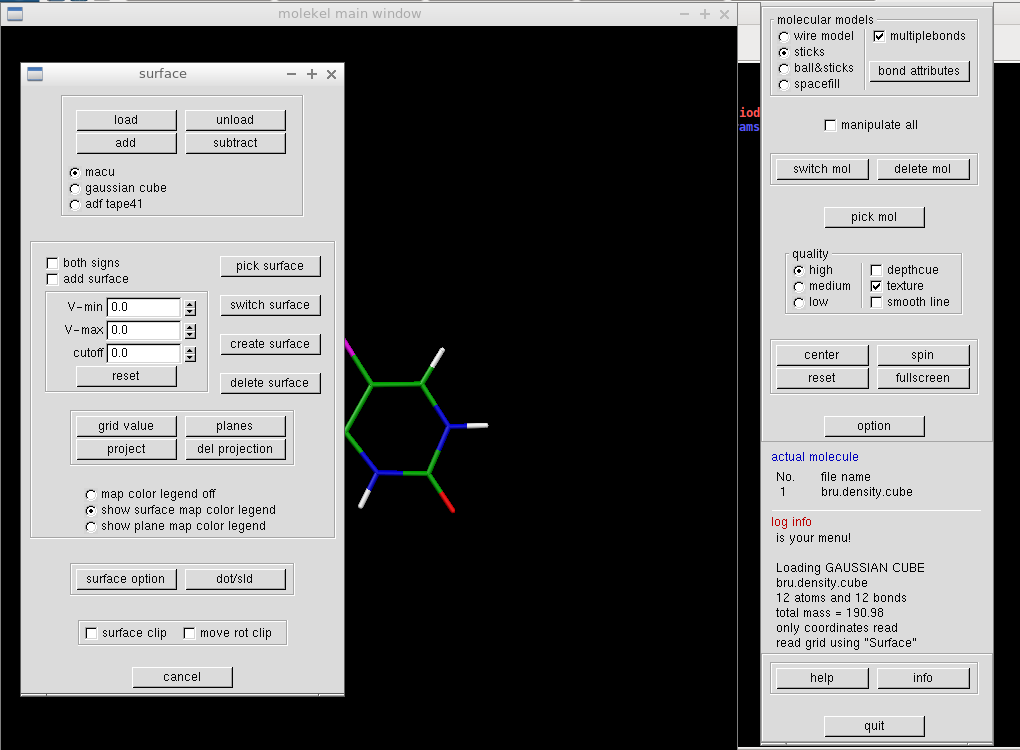
We construct the density isosurface by specifying a density value in "cutoff" field and by clicking "create surface" button. A white molecular surface should appear on the main screen.
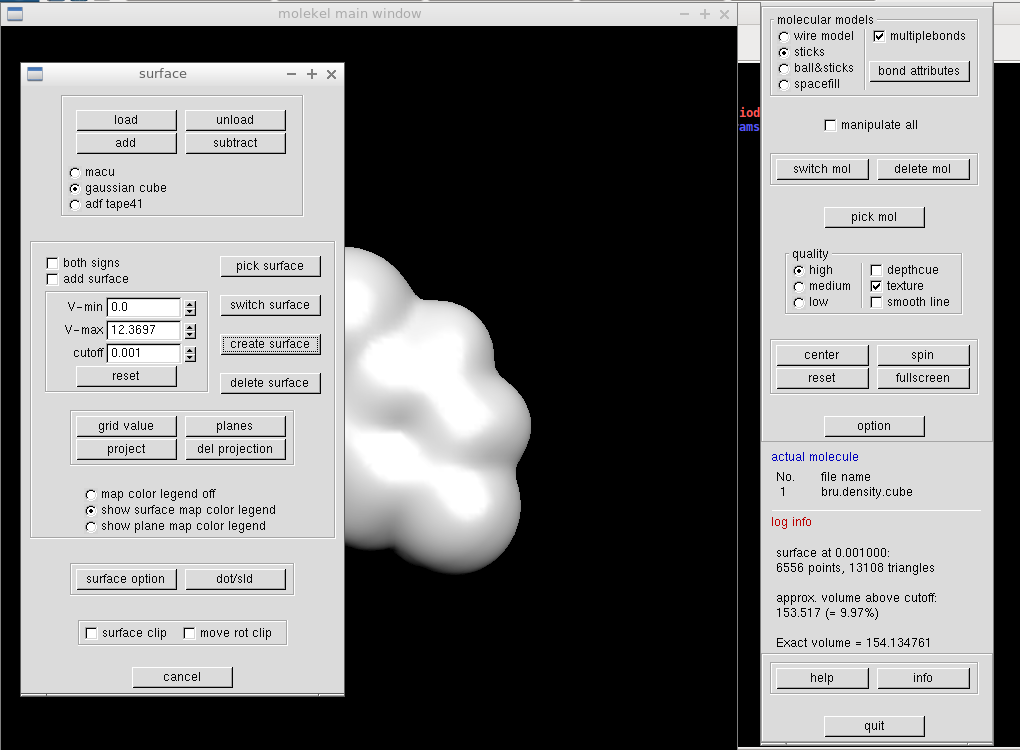
We want to color the surface according to the electrostatic potential. In the same "Surface" window, we click the "load" button and select the cube file with the electrostatic potential (here mol.pote.cube). The V-min and V-max values should change. Clicking on "grid value" button makes the surface colorful and opens a new window. It is reasonable to set the minimum and maximum values identical (of course apart from their signs).
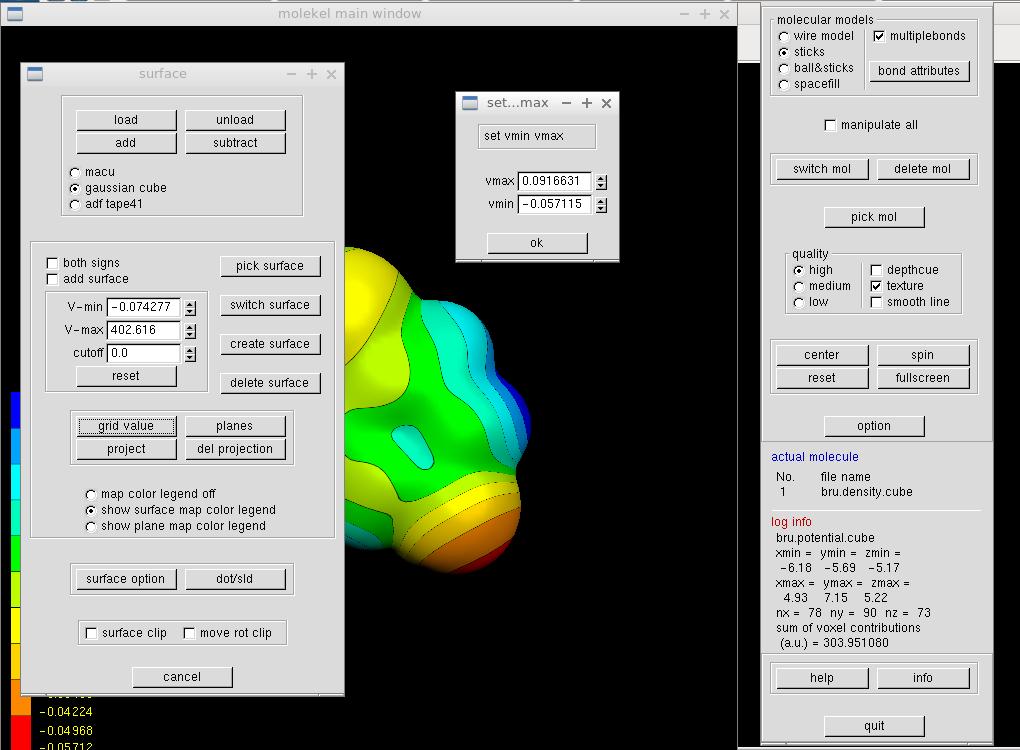
We may wish to adjust the coloring scheme. Such a change can be done in the "Texture" window from the main menu (right click in the main window with the molecule and its colored surface). The buttons list up and down offer a few variants. One adds the texture to the surface by clicking the "surface" button in the "Texture" window, and the molecular surface in the main window immediately after.
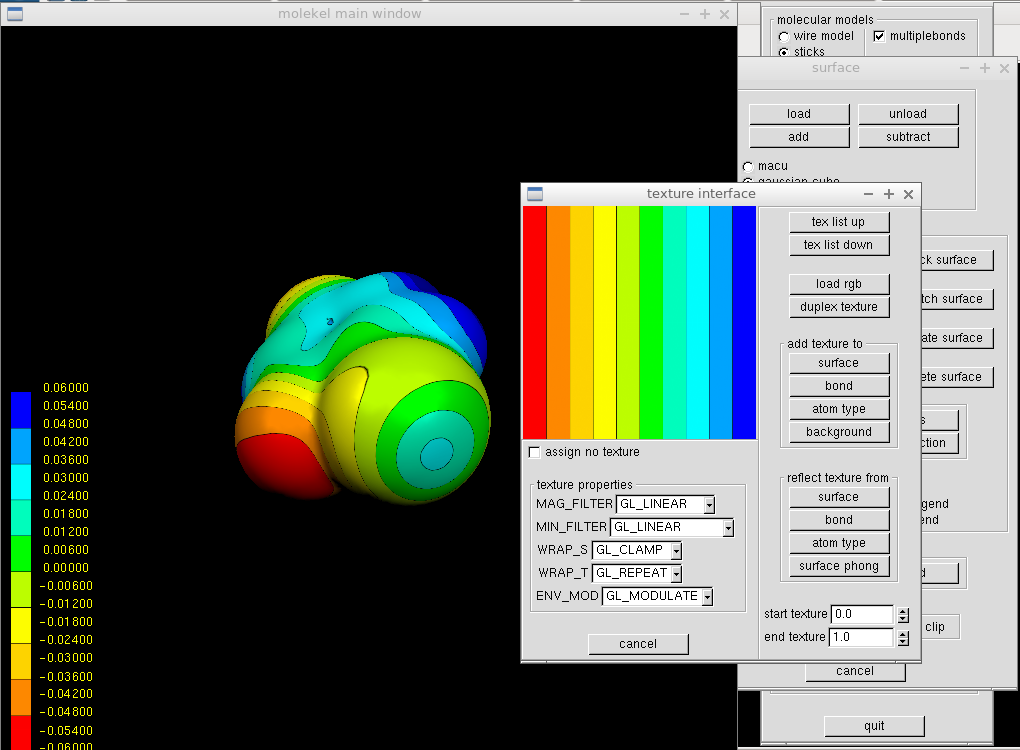
The background can be changed in the "Color" window from the main menu. For a white background, we put 1.0 for all diffuse color fields and click "top" and "bottom." Similarly, one can adjust label color to black.
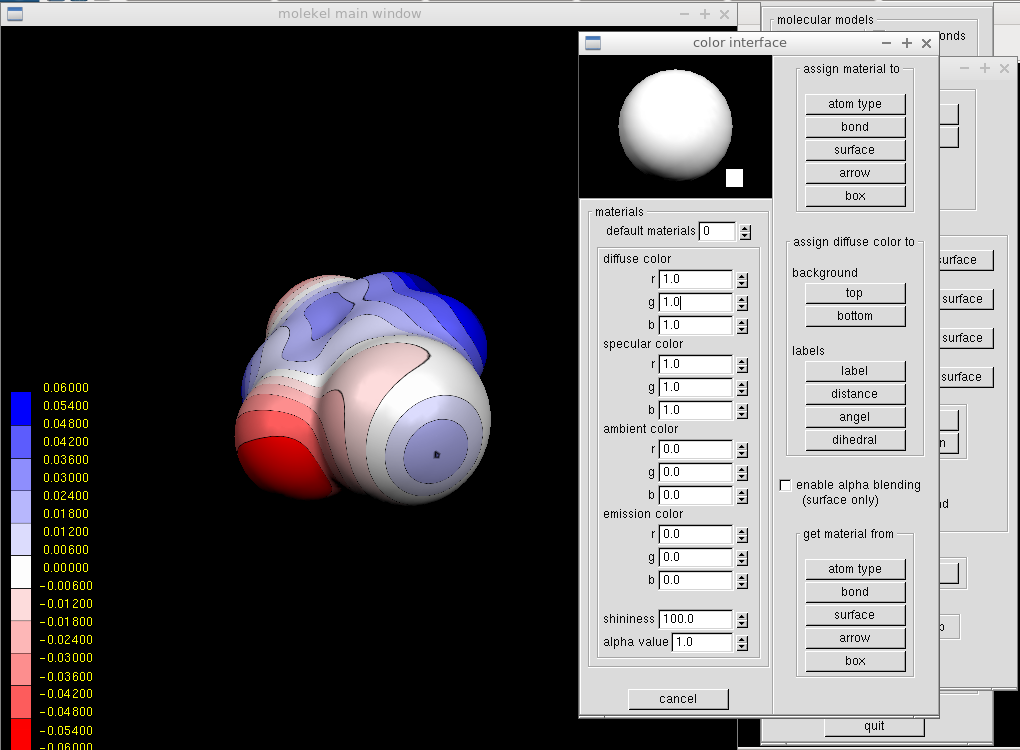
It remains to render the image by "Render" from the main menu. We set large dimensions and jpeg file type with 100 % quality. After specifying the file name, we can enjoy an eye-catching molecular electrostatic potential and adjust it further in any bitmap image processor.
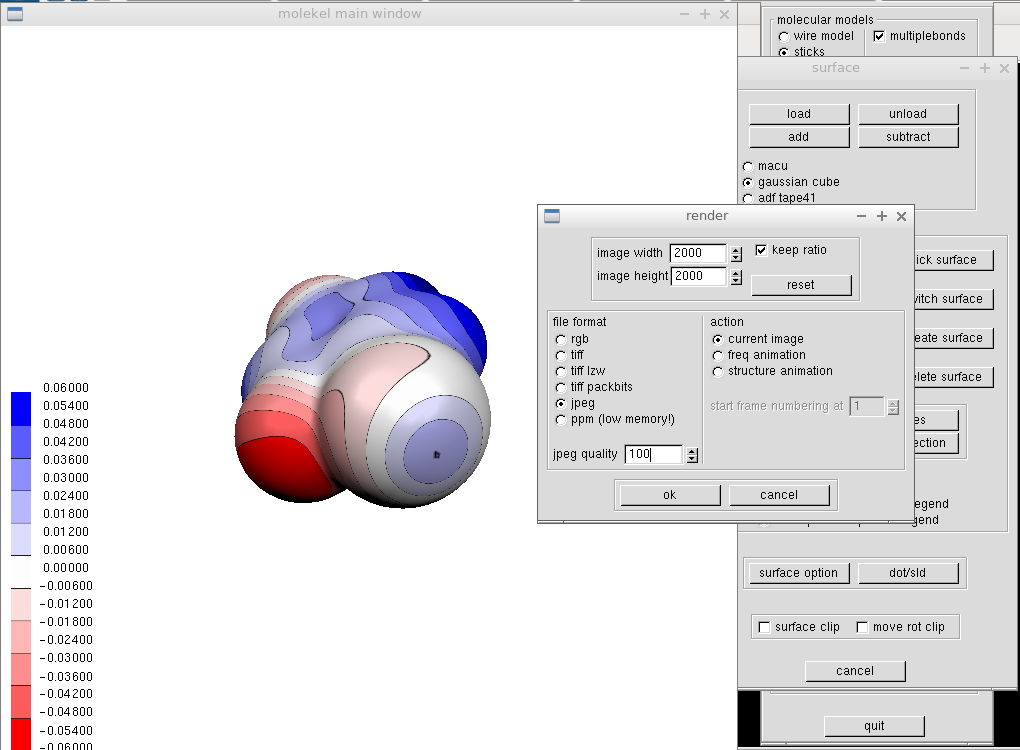
I usually use Gimp and Inkscape. This way, I prepared MEPs for several publications on halogen bonding. See e.g.
- 2017
- Halogen Bonding in Nucleic Acid Complexes
- 2016
- Computer Modeling of Halogen Bonds and Other sigma-Hole Interactions
- 2013
- The differences in the sublimation energy of benzene and hexahalogenbenzenes are caused by dispersion energy
Was it useful? Are there any errors? Drop me a line to
 .
I'll be happy to see your production...
.
I'll be happy to see your production...
