Two bachelor theses defended with honors
published 2021-08-29
Two students from our group obtained their BSc. with honors. Warmest congratulations!
Hugo defended his thesis about MD simulations of the ribosome, where he focused on the interaction between ribosome surface and an enzyme important for protein biogenesis. Hugo plans to stay with the group pursuing his interest in simulations and advanced statistical analysis (aka machine learnine).
Iva defended her thesis about nascent protein unfolding within the ribosome tunnel. She used enhanced sampling techniques to quantify energetics of co-translational folding/unfolding. From September 2021, she's a student of Neurobiology at University of Copenhagen.
Protein birth on the Summer school for teachers
published 2021-08-26
Michal gave a talk on the 35th Summer school for high-school teachers and students. The event was organized by UCT Prague and the talks took place in the Balling hall of the National library of technology. The modern lecture hall is named after a Prague chemist, who, among other achievements, developed a scale for beer strength.
The talk was entitled "How proteins are born". Michal usually prefers slides with low information content. Anyways, for Czech audience they are available here. If you wish to put yourself in the protein mood, give yourself a bisceps massage or ruffle your hair. They are full of protein.
Arian ranked 5th in a student research competition
published 2021-06-25
Arian Adam Ott is a high-school student from a small town in West Bohemia. Last year, he started studying short peptides in the ribosome exit tunnel using atomistic simulations. Now he ranked 5th in Biology section in the National exhibition of student research projects, also known as SOČ. Congratulations! We are looking forward to your future studies at UCT Prague.

Michal received Humboldt Alumni Award 2021
published 2021-05-07
Pleased to inform that this year Humboldt Alumni Award goes to Michal H. Kolář from UCT Prague. The Alexander von Humboldt Foundation is a German organization, which promotes international academic cooperation between excellent scientists and scholars. Each year, the Foundation awards up to 5 individuals from all over the world, who promote novel networking initiatives. Michal succeeded with a project with Czexpats in Science about sharing the academic experience of Czech researchers working abroad with those who reside in Czechia. The award will be officially announced during the virtual Annual Meeting in June.

The not so short story about writing a grant proposal
last update 2021-03-27
I have recently lost a collaborator with which we were preparing a grant proposal. He changed his mind and went with another more senior/competitive/secured partner. Because there was not a place for me in their proposal, I decided to apply on my own. Unfortunately, I must start from scratch because the previous topic does not make sense without the collaboration. So I take this opportunity to share my thoughts about the grant application, and write a cookbook which may be useful especially for those sailing in the deep waters of the Czech academic environment. The story might not be short (Latex users know).
The deadline is April 8. I'm starting now.

Three grants of the Internal Grant Agency approved
published 2021-03-01
UCT Prague offers one-year funding through its Internal Grant Agency. We are pleased to announced that from our lab three grants, one research and two teaching, were approved. The research grant goes to Felipe, who will study peptide behavior in carbon nanotube. Michal is a PI and co-PI of two teaching grants, which aim at improving physical chemistry courses via an online search tool. Stay tuned for the updates.

Supervision workshop in Brno
published 2020-10-10
Michal co-organized a workshop for PhD supervisors at Masaryk University Brno (MUNI). It was a second round of workshops organized by Czexpats in Science and the Rector's office for early career researchers and group leaders. Apart from Michal, who gave a talk about supervisor-student relationship, three speakers shared their experience: Monika Čechová from Penn State, USA, Tomáš Kubař from Karsruhe Institute of Technology, Germany, and Olga Löblová from Cambridge University, UK. Michal's slides are available here (in Czech).
První víkend s Linuxem
The first weekend with Linux
last update 2020-08-31
Undergraduate students often come to me with their interest in computational chemistry and/or biomolecular dynamics. It is a fascinating research area indeed, but I somewhat feel that the beginning is either hard or slow. So far, I haven't found an effective way to start investigating molecules without a decent knowledge of bash, python, and some specific computational chemistry program. I know there are many tutorials with toy models with known output to reproduce, etc., but I find such work less attractive than pure exploration. Hence I wrote this Linux intro suited specifically for what we do in my group at UCT. To decrease the activation barrier even more, it's in Czech. I hope you'll find English resources, should your Czech be poorer than bash.
Aneta and Ondra defended their theses
published 2020-07-10
Two members of our team have defended their theses and successfully finished bachelor studies. They've became the first students obtaining a degree under Michal's supervision. Ondra is leaving to study proteins experimentally at the Department of microbiology at UCT. Aneta is leaving as well, but we hope it will be just temporarilly. She plans to discover the student's life in Portugal during an Erasmus stay. Best wishes to both of them.
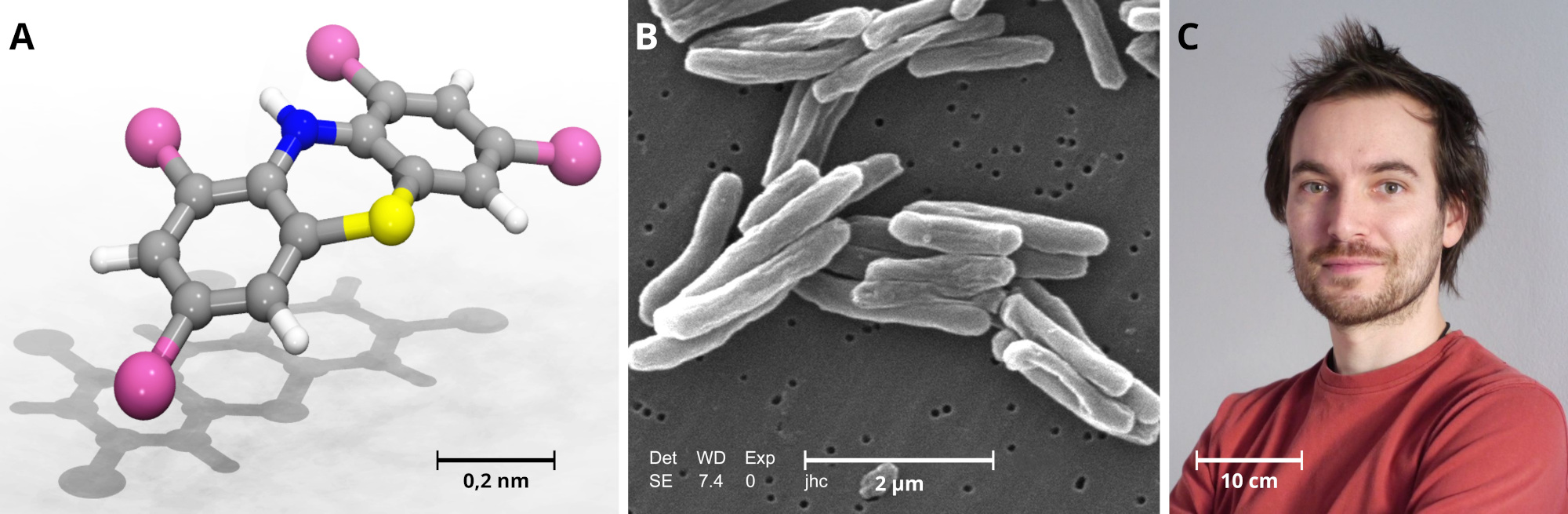
Polyhalogenated phenothiazines to fight tuberculosis
published 2020-06-30
A press release about our recent paper.
The Mycobacterium tuberculosis causes over a million deaths annually. Especially in third-world countries, it is still a severe problem, which may even worsen due to antibiotic resistance. An international team of researchers, together with Dr. Michal H. Kolář from UCT Prague, has now developed new compounds to fight the bacterium, which had cost George Orwell or Jane Austen their lives.